Clustering by fitting a mixture model using EM with K groups
and unconstrained covariance matrices for a matrix variate normal or
matrix variate t distribution (with specified degrees of freedom nu).
matrixmixture(
x,
init = NULL,
prior = NULL,
K = length(prior),
iter = 1000,
model = "normal",
method = NULL,
row.mean = FALSE,
col.mean = FALSE,
tolerance = 0.1,
nu = NULL,
...,
verbose = 0,
miniter = 5,
convergence = TRUE
)Arguments
- x
data, \(p \times q \times n\) array
- init
a list containing an array of
Kof \(p \times q\) means labeledcenters, and optionally \(p \times p\) and \(q \times q\) positive definite variance matrices labeledUandV. By default, those are presumed to be identity if not provided. Ifinitis missing, it will be provided using thepriororKbyinit_matrixmix.- prior
prior for the
Kclasses, a vector that adds to unity- K
number of classes - provide either this or the prior. If this is provided, the prior will be of uniform distribution among the classes.
- iter
maximum number of iterations.
- model
whether to use the
normalortdistribution.- method
what method to use to fit the distribution. Currently no options.
- row.mean
By default,
FALSE. IfTRUE, will fit a common mean within each row. If both this andcol.meanareTRUE, there will be a common mean for the entire matrix.- col.mean
By default,
FALSE. IfTRUE, will fit a common mean within each row. If both this androw.meanareTRUE, there will be a common mean for the entire matrix.- tolerance
convergence criterion, using Aitken acceleration of the log-likelihood by default.
- nu
degrees of freedom parameter. Can be a vector of length
K.- ...
pass additional arguments to
MLmatrixnormorMLmatrixt- verbose
whether to print diagnostic output, by default
0. Higher numbers output more results.- miniter
minimum number of iterations
- convergence
By default,
TRUE, using Aitken acceleration to determine convergence. If false, it instead checks if the change in log-likelihood is less thantolerance. Aitken acceleration may prematurely end in the first few steps, so you may wish to setminiteror selectFALSEif this is an issue.
Value
A list of class MixMatrixModel containing the following
components:
priorthe prior probabilities used.
initthe initialization used.
Kthe number of groups
Nthe number of observations
centersthe group means.
Uthe between-row covariance matrices
Vthe between-column covariance matrix
posteriorthe posterior probabilities for each observation
pithe final proportions
nuThe degrees of freedom parameter if the t distribution was used.
convergencewhether the model converged
logLika vector of the log-likelihoods of each iteration ending in the final log-likelihood of the model
modelthe model used
methodthe method used
callThe (matched) function call.
References
Andrews, Jeffrey L., Paul D. McNicholas, and Sanjeena Subedi. 2011.
"Model-Based Classification via Mixtures of Multivariate
T-Distributions." Computational Statistics & Data Analysis 55 (1):
520–29. \doi{10.1016/j.csda.2010.05.019}.
Fraley, Chris, and Adrian E Raftery. 2002. "Model-Based Clustering,
Discriminant Analysis, and Density Estimation." Journal of the
American Statistical Association 97 (458). Taylor & Francis: 611–31.
\doi{10.1198/016214502760047131}.
McLachlan, Geoffrey J, Sharon X Lee, and Suren I Rathnayake. 2019.
"Finite Mixture Models." Annual Review of Statistics and Its
Application 6. Annual Reviews: 355–78.
\doi{10.1146/annurev-statistics-031017-100325}.
Viroli, Cinzia. 2011. "Finite Mixtures of Matrix Normal Distributions
for Classifying Three-Way Data." Statistics and Computing 21 (4):
511–22. \doi{10.1007/s11222-010-9188-x}.See also
Examples
set.seed(20180221)
A <- rmatrixt(20,mean=matrix(0,nrow=3,ncol=4), df = 5)
# 3x4 matrices with mean 0
B <- rmatrixt(20,mean=matrix(1,nrow=3,ncol=4), df = 5)
# 3x4 matrices with mean 1
C <- array(c(A,B), dim=c(3,4,40)) # combine into one array
prior <- c(.5,.5) # equal probability prior
# create an intialization object, starts at the true parameters
init = list(centers = array(c(rep(0,12),rep(1,12)), dim = c(3,4,2)),
U = array(c(diag(3), diag(3)), dim = c(3,3,2))*20,
V = array(c(diag(4), diag(4)), dim = c(4,4,2))
)
# fit model
res<-matrixmixture(C, init = init, prior = prior, nu = 5,
model = "t", tolerance = 1e-3, convergence = FALSE)
print(res$centers) # the final centers
#> , , 1
#>
#> [,1] [,2] [,3] [,4]
#> [1,] 0.117537755 0.05142832 0.03229732 -0.02163010
#> [2,] 0.005705083 -0.03489656 -0.03159498 -0.05027392
#> [3,] -0.041192795 -0.10381233 -0.07923590 -0.06302149
#>
#> , , 2
#>
#> [,1] [,2] [,3] [,4]
#> [1,] 1.065892 1.0573621 0.8615166 1.0380062
#> [2,] 1.069079 0.9432203 1.0902007 0.9915732
#> [3,] 1.080176 1.0671593 0.9880986 0.7968937
#>
print(res$pi) # the final mixing proportion
#> [1] 0.4987695 0.5012305
plot(res) # the log likelihood by iteration
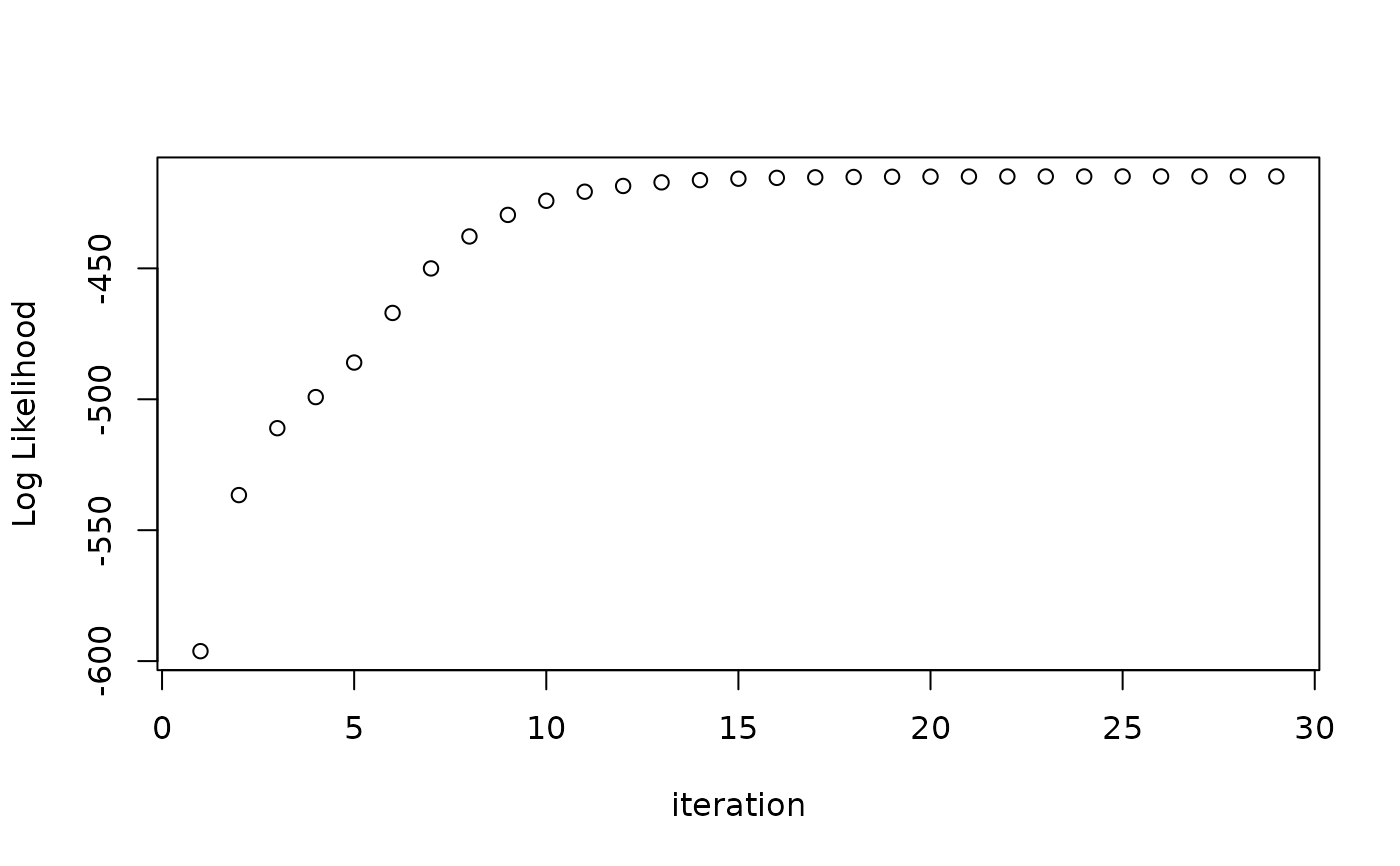 logLik(res) # log likelihood of final result
#> 'log Lik.' -414.8875 (df=54)
BIC(res) # BIC of final result
#> [1] 1028.975
predict(res, newdata = C[,,c(1,21)]) # predicted class membership
#> $class
#> [1] 1 2
#>
#> $posterior
#> [,1] [,2]
#> [1,] 9.999883e-01 1.170306e-05
#> [2,] 6.703426e-07 9.999993e-01
#>
logLik(res) # log likelihood of final result
#> 'log Lik.' -414.8875 (df=54)
BIC(res) # BIC of final result
#> [1] 1028.975
predict(res, newdata = C[,,c(1,21)]) # predicted class membership
#> $class
#> [1] 1 2
#>
#> $posterior
#> [,1] [,2]
#> [1,] 9.999883e-01 1.170306e-05
#> [2,] 6.703426e-07 9.999993e-01
#>